Using evolutionary theory and genomic tools to enhance conservation
Distribution of Diversity
|
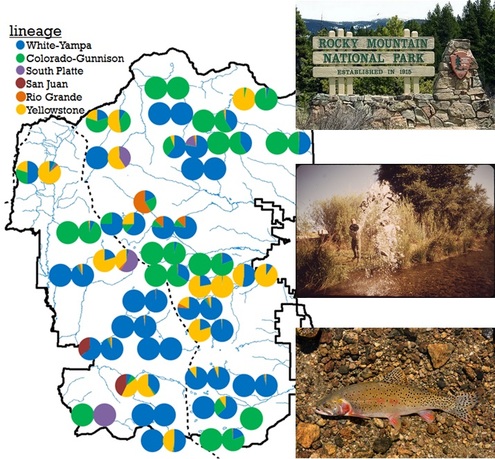
The foundation of both evolutionary biology and conservation is the accurate description of the biological unit. My work on bighorn sheep and cutthroat trout starts with the detection of biological clusters based on shared genotypes and identifying where that diversity is found. This is accomplished using population genetic clustering and phylogenetic methods using tens to tens of thousands of genetic markers. Using complementary marker types from microsatellites to mitochondrial sequences to genomic SNPs allows me to access information at different scales: from fine-scale at the levels of individuals, families, and drainages, and recent history within only a few generations; to large-scale at the levels of evolutionary lineages, species, ranges, and thousands of generations. Both levels of information are necessary for understanding evolutionary processes and for effective conservation.
|
How is the history of human intervention reflected in the genomes of individuals? I answer this question by looking at the distribution of genetic diversity across the landscape and connecting that information from the historical record. Both cutthroat trout and bighorn sheep have extensive stocking and translocation records that help us understand how evolutionarily distant genomes end up in close geographic proximity or even mixed in the same individual.
Manipulation of Genomes
Diminishing numbers and isolation from other populations can lead to a loss of genetic diversity and fitness due to inbreeding and the accumulation of deleterious alleles. Using a combination of molecular genetic techniques and controlled and natural experiments, I investigate what measures will be most effective for increasing genetic diversity and recovering demographic stability. Human actions have inadvertently altered the genomes of species such as cutthroat trout for a variety of purposes, but intentionally manipulating their genomes may become an important conservation strategy.
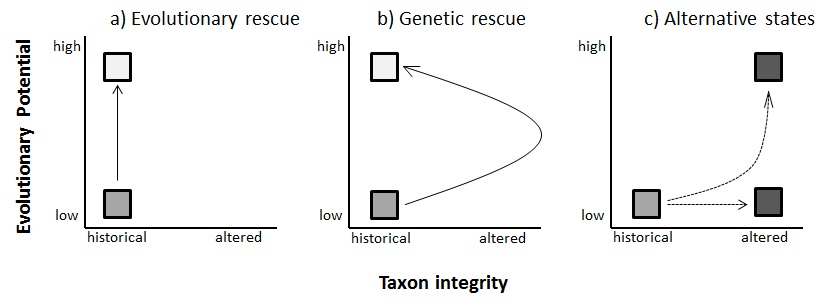
Drawing on the field of ecological restoration, my work is guided by a model of evolutionary restoration in which the restoration of genetic diversity and fitness is visualized as a relationship between taxonomic integrity and evolutionary potential. There are two general paths by which we can recover evolutionary potential, the individual fitness and population diversity needed to respond to selection. First, a population may return to high evolutionary potential without disrupting taxonomic integrity, such as by breeding designs within a subspecies. The second pathway involves temporarily disrupting taxonomic integrity by intentionally outcrossing the target population with another population, subspecies, or even species. Of course, this strategy runs the risk of resulting in alternative states, such as outbreeding depression or hybrid swarms. For small inbred populations that are the sole remaining representatives of the species and are trapped in extinction vortices, genetic rescue may be the only option to prevent extinction.
|
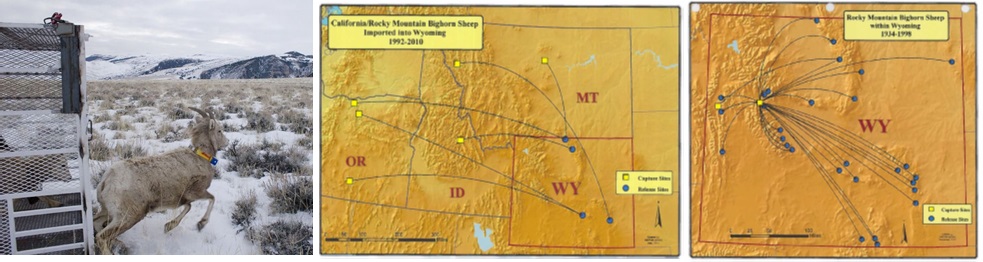
I used controlled crosses in cutthroat trout to test these two alternative strategies. I investigated the role of relatedness in offspring fitness, looking at a range of relatedness from parents who are siblings to parents from different species. I am interested in extending this work to examine the role of the number and identity of introduced individuals in genetic rescue. I plan to use natural experiments such as the translocation of bighorn sheep into existing populations and tractable mesocosm experiments with small fishes.
|
|